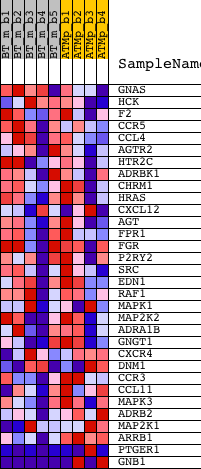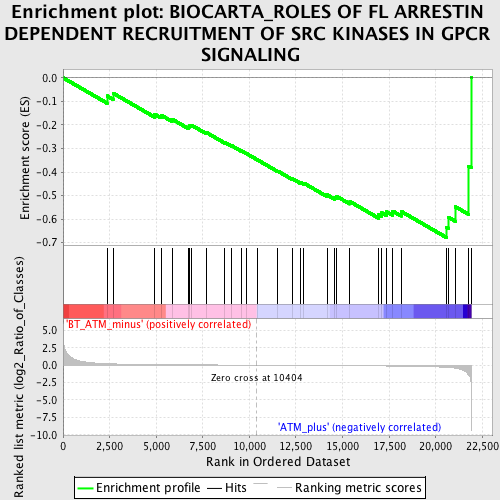
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_BT_ATM_minus_versus_ATM_plus.cls#BT_ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
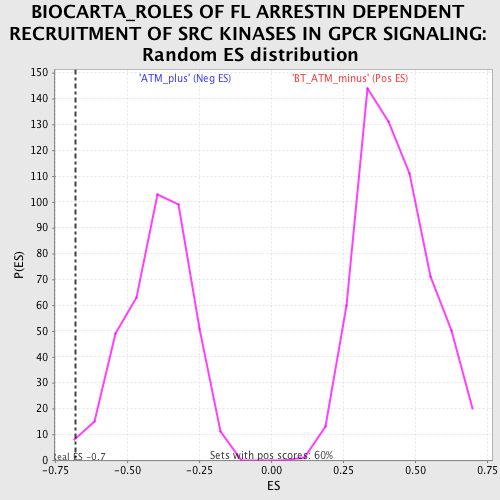
| GeneSet | BIOCARTA_ROLES OF FL ARRESTIN DEPENDENT RECRUITMENT OF SRC KINASES IN GPCR SIGNALING |
| Enrichment Score (ES) | -0.6779047 |
| Normalized Enrichment Score (NES) | -1.725645 |
| Nominal p-value | 0.015037594 |
| FDR q-value | 0.23175049 |
| FWER p-Value | 0.933 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GNAS | 1421740_at 1427789_s_at 1443007_at 1443375_at 1444767_at 1450186_s_at 1453413_at | 2363 | 0.216 | -0.0770 | No | ||
| 2 | HCK | 1446553_at 1449455_at | 2686 | 0.183 | -0.0655 | No | ||
| 3 | F2 | 1418897_at | 4917 | 0.089 | -0.1546 | No | ||
| 4 | CCR5 | 1422259_a_at 1422260_x_at 1424727_at | 5269 | 0.081 | -0.1590 | No | ||
| 5 | CCL4 | 1421578_at | 5852 | 0.069 | -0.1758 | No | ||
| 6 | AGTR2 | 1415832_at | 6741 | 0.053 | -0.2088 | No | ||
| 7 | HTR2C | 1435513_at 1450477_at | 6766 | 0.052 | -0.2024 | No | ||
| 8 | ADRBK1 | 1426249_at 1451992_at | 6880 | 0.050 | -0.2004 | No | ||
| 9 | CHRM1 | 1439611_at 1450833_at | 7689 | 0.037 | -0.2320 | No | ||
| 10 | HRAS | 1422407_s_at 1424132_at | 8692 | 0.023 | -0.2744 | No | ||
| 11 | CXCL12 | 1417574_at 1439084_at 1448823_at | 9049 | 0.018 | -0.2881 | No | ||
| 12 | AGT | 1423396_at | 9556 | 0.011 | -0.3096 | No | ||
| 13 | FPR1 | 1450808_at | 9828 | 0.008 | -0.3208 | No | ||
| 14 | FGR | 1419526_at 1442804_at | 10416 | -0.000 | -0.3476 | No | ||
| 15 | P2RY2 | 1450318_a_at | 11525 | -0.015 | -0.3961 | No | ||
| 16 | SRC | 1423240_at 1450918_s_at | 12329 | -0.025 | -0.4292 | No | ||
| 17 | EDN1 | 1451924_a_at | 12748 | -0.031 | -0.4439 | No | ||
| 18 | RAF1 | 1416078_s_at 1425419_a_at | 12932 | -0.033 | -0.4475 | No | ||
| 19 | MAPK1 | 1419568_at 1426585_s_at 1442876_at 1453104_at | 14178 | -0.052 | -0.4970 | No | ||
| 20 | MAP2K2 | 1415974_at 1443436_at 1460636_at | 14594 | -0.058 | -0.5076 | No | ||
| 21 | ADRA1B | 1422183_a_at | 14696 | -0.060 | -0.5036 | No | ||
| 22 | GNGT1 | 1425167_a_at 1425168_at 1425219_x_at 1451633_a_at | 15403 | -0.072 | -0.5256 | No | ||
| 23 | CXCR4 | 1448710_at | 16946 | -0.103 | -0.5813 | No | ||
| 24 | DNM1 | 1427754_a_at 1456346_at 1460365_a_at | 17114 | -0.108 | -0.5735 | No | ||
| 25 | CCR3 | 1422957_at | 17364 | -0.114 | -0.5685 | No | ||
| 26 | CCL11 | 1417789_at | 17688 | -0.123 | -0.5656 | No | ||
| 27 | MAPK3 | 1427060_at | 18157 | -0.138 | -0.5673 | No | ||
| 28 | ADRB2 | 1437302_at | 20580 | -0.309 | -0.6337 | Yes | ||
| 29 | MAP2K1 | 1416351_at | 20684 | -0.326 | -0.5918 | Yes | ||
| 30 | ARRB1 | 1460444_at | 21076 | -0.432 | -0.5480 | Yes | ||
| 31 | PTGER1 | 1436542_at 1445445_s_at 1450491_at | 21785 | -1.439 | -0.3748 | Yes | ||
| 32 | GNB1 | 1417432_a_at 1425908_at 1454696_at 1459442_at | 21915 | -2.671 | 0.0009 | Yes |